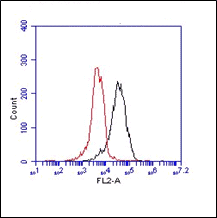
Antibodies derived from clone MaP.mGILT6 recognize mouse gamma-interferon-inducible lysosomal thiol reductase (GILT). GILT facilitates major histocompatibility complex (MHC) class II-restricted processing through reduction of protein disulfide bonds in the endocytic pathway. GILT can also facilitate the transfer of disulfide-containing antigens into the cytosol, enhancing their cross-presentation by MHC class I.
Human GILT consists of 261 amino acids with a 37 amino acid signal sequence and a 224 amino acid precursor form. In general, the 35 kDa precursor is tagged with mannose-6- phosphate (M6P) residues and transported to the endocytic pathway via the M6P receptor (M6PR). A small fraction of the precursor is secreted as a disulfide-linked dimer. In early endosomes, N- and C-terminal pro-peptides are cleaved to generate a 28 kDa mature form. The mature form of GILT is localized to late endosomes and lysosomes and has maximal reductase activity at the acidic pH found in these compartments.
GILT is constitutively expressed in professional antigen presenting cells (APC). The constitutive expression of GILT in APCs is likely to account for enhanced reduction of proteins in the late endosomal, lysosomal, and phagosomal compartments. A combination of GILT-mediated reduction and proteolysis in the these compartments can enhance the generation of class II epitopes as well as facilitate the translocation of internalized antigens into the cytosol for proteosomal processing and cross-presentation. It is also constitutively expressed in thymocytes, mature T cells, and some fibroblasts. IFN-ɣ plays is the key factor in inducing expression of MHC class II and other components of the class II-restricted processing pathway. IFN-ɣ signals through a specific cell surface receptor followed by activation of JAK/STAT1 signalling.